 |
 |
- Search
| J Korean Med Assoc > Volume 57(3); 2014 > Article |
Abstract
Atopic dermatitis (AD) is a chronic relapsing inflammatory skin disease with severe pruritus, and the first step of atopic march since it often precedes asthma or allergic rhinitis. Since its etiology or pathogenesis is very complex and frequently changing, physicians cannot easily understand it in entirety. New insights into the genetics and pathophysiology of AD emphasize the crucial function of the skin barrier as well as abnormal immune response. In this review, the pathogenesis of AD is explained as the combined features of impaired skin barrier and abnormal immune response rather than each independent concept. Understanding the whole pathogenesis of AD may lead to early intervention and prevention of atopic march as well as proper clinical treatment.
아토피피부염은 산업이 발달하고 핵가족화가 이루어지는 서구화된 나라에서 더욱 흔하게 나타나는 질환으로 소아는 10-20%, 성인은 1-3% 정도의 유병률을 보이는데 최근 급격한 증가추세를 보이는 우리나라의 경우도 이와 유사한 유병률을 나타내고 있다[1]. 아토피피부염 환자의 70-80% 정도에서 아토피질환의 가족력이 있을 만큼 아토피피부염은 유전적인 소인이 매우 중요하게 작용하지만, 우리나라처럼 급격한 서구화 및 산업화와 함께 아토피피부염의 발생이 현저하게 증가하는 것은 환경적인 요인도 강력하게 영향을 미치고 있음을 나타낸다. 이를 위생가설(hygiene hypothesis)로 설명하기도 하지만 완벽하지는 않다. 2006년 필라그린 유전자(FLG)의 기능소실 돌연변이(loss of function mutation)가 북서유럽국가의 백인 아토피피부염 환자에서 매우 중요하게 관여함이 보고된 이후에는 선천적인 피부장벽 손상이 전통적으로 중요시되던 비정상적인 알레르기 반응을 대신하여 아토피피부염의 1차적인 원인으로도 대두되었다[2]. 최근에는 아토피피부염의 병인을 피부장벽 이상이 1차적으로 작용한다는 outside-in model과 비정상적인 면역반응이 1차적으로 발생하여 시작된다는 inside-out model로 크게 두 가지로 나누고 있으나, 이 두 기전이 상호복합적으로 작용한다는 outside-inside-outside model로 통합적으로 설명하고 있다[3]. 즉, 취약한 피부장벽과 비정상적인 면역반응이 합해져서 아토피피부염의 병인에 기여한다. 따라서 본 종설에서는 아토피피부염의 병인을 유전학적인 측면까지 포함하여 크게 피부장벽의 기능이상 및 비정상적인 면역반응으로 나누어 기술한다.
피부장벽 기능이상(skin barrier dysfunction)과 연관되어 제시되는 병인들[4] 중 대표적인 것으로 필라그린 유전자(FLG) 돌연변이[5], 각질층의 세라마이드 감소[6], 항균 펩타이드 감소[7], 세린계단백질분해효소 억제제(serine proteases inhibitor) 감소[8], 밀찹연접(tight junction) 이상[9] 등이 있다.
아토피피부염 환자의 비병변 피부에서 표피분화 단백질 중 유독 필라그린의 발현만이 감소되었다는 결과가 보고된 후[10] 아토피피부염의 병인으로 필라그린을 주목하는 여러 연구가 진행되었고, 현재는 유럽 및 아시아 등지에서 총 27가지 이상이 FLG 기능소실 돌연변이로 보고되었다. 북서유럽 백인의 아토피피부염 환자 중 FLG 변이는 40% 정도에서 나타나고, 중국인은 약 30%, 일본인은 약 25%, 한국인은 8% 미만에서 보고되고 있으며[11], FLG 변이 중 R501X와 E2422X만이 북서유럽 백인과 아시아인에서 동시에 보고 되었을 정도로 FLG 변이는 인종에 따른 뚜렷한 차이를 보이고 있다[12,13,14,15]. FLG 변이가 있는 아토피피부염 환자들은 더욱 어린 나이에 아토피피부염이 발병하며, 더욱 심한 증상을 보이면서, 성인 시기까지 지속되는 경우가 더 많고, 호흡기 아토피질환으로 진행하는 알레르기행진을 보이는 경우가 더 많다는 것이 밝혀졌다[16]. 필라그린은 표피 과립층에서 케라토히알린과립(keratohyalin granule)을 형성하는 단백질인 프로필라그린(profilaggrin)의 형태로 존재하다가, 각질형성세포의 최종분화과정에서 필라그린으로 분해된 후에 각질세포막을 형성할 때 케라틴 필라멘트를 응집하여 각질세포의 단단하고 편평한 구조를 만듦으로써 피부장벽에서 벽돌(brick)의 역할을 수행하게 된다. 이후 필라그린은 각질층 상부로 올라가면서 아미노산으로 분해되어 자유아미노산, pyrrolidone carboxylic acid (PCA), urocanic acid (UCA), 다양한 유기산 등으로 변화하게 된다. 자연보습인자(natural moisturizing factor) 중에서 자유아미노산이 40%, PCA가 12%를 차지하며, 이들 필라그린 분해산물은 각질층의 수화뿐 아니라 각질층 pH의 정상화, 투과장벽, 각질층의 견고함(integrity/cohesion), 항균 및 항염작용 등에서 다양한 역할을 수행한다(Figure 1) [4]. 즉 FLG 변이가 발생하면 필라그린 단백질 생성이 감소되어 각질세포막 형성을 약화시키고, 각질세포 사이의 접착력을 감소시키며, 경표피수분손실(transepidermal water loss)이 증가됨으로써, 전반적인 피부장벽기능이 감소되어 외부 알레르겐의 체내 침투 증가에 의한 감작 및 이로 인한 알레르기 반응이 쉽게 일어난다. 또한 세포 골격이 위축되기 때문에 층판소체의 내용물이 부실해지고, trans-UCA 감소에 의해서 각질층 pH가 증가하며, 분해산물인 아미노산과 PCA가 감소하기 때문에 건조피부가 유발됨으로써 아토피피부염이 발병하게 된다[17].
아토피피부염 환자의 병변뿐만 아니라 건조피부만을 보이는 비병변 피부에서도 각질세포간 지질막을 형성하는 주성분인 세라마이드(ceramides)가 뚜렷하게 감소하고, 특히 가장 긴 사슬구조인 세라마이드 1이 가장 많이 감소함을 보여서, 세라마이드의 감소를 필라그린 분해산물인 자연보습인자의 감소와 함께 거의 모든 아토피피부염 환자에서 관찰되는 건조피부의 중요한 원인으로 생각하였다[18] (Figure 2). 아토피피부염 환자의 피부는 항균 펩타이드(antimicrobial peptide)가 감소되어있고 특히 각질층에서 세라마이드 분해산물이며 가장 강력한 항균작용을 하는 스핑고신(sphingosine)의 생성이 감소되어 있어 세균, 바이러스, 진균 등의 감염에 취약하게 된다[7].
아토피피부염 환자의 피부표면 pH는 정상인보다 높은데, 각질층 pH가 증가하면 각질세포의 탈락을 유도하는 세린계단백질분해효소(serine protease)의 활성이 촉진되고 protease-activated receptor-2 (PAR-2)의 활성이 증가되어 Th2 면역반응이 우선하는 알레르기 염증반응이 일어나게 된다. 아토피피부염 및 천식의 주요한 원인항원인 집먼지 진드기, 바퀴벌레 항원 등은 PAR-2를 활성화시킴으로써 손상된 피부장벽의 회복을 지연시키고 아토피피부염을 악화시킨다는 것이 확인 되었다[19]. 그런데 이 분해효소의 작용을 억제시키는 역할을 수행하는 단백질이 LEKTI (lympho-epithelial Kazal type-related inhibitor)로 serine protease inhibitor Kazal type 5 유전자(SPINK5)의 산물이다. 아토피피부염 환자 중 염색체 5q32에 위치한 SPINK5 변이가 보고되었고[8], 이들 환자들에서는 활성화된 세린계단백질분해효소의 작용을 적절하게 억제하지 못하여 알레르기 염증반응이 증가하는 것으로 설명할 수 있다(Figure 3). 한편으로는 세린계단백질분해효소인 Kallikrein7 (stratum corneum chymotryptic enzyme) 유전자(KLK7) 변이와 아토피피부염과의 관련성도 보고되었다[20].
밀착연접(tight junction)은 상피세포에 존재하며 이웃한 세포막 측면을 서로 고정시키고 세포 측면 공간을 통한 수분과 체액의 흐름을 방지하는 장벽이다. 각질층이 손상되면서 각질층과 과립층 사이에 존재하는 체액의 경표피수분손실과 함께 칼슘이온 농도도 감소하면 바로 아래의 과립층 상부의 밀착연접이 열리고[21], 밀착연접 바로 아래까지 위치하고 있던 랑게르한스세포의 수지(dendrite)가 밀착연접 위로 올라오게 된다. 이때 손상된 각질층을 통과하여 들어온 분자량이 큰 단백질 항원들은 랑게르한스세포의 수지에 의해 포집되고 인식되어 국소 림프절로 이동하고, 이들 단백질 항원의 세린계단백분해효소 성분은 각질형성세포막에 존재하는 PAR-2를 활성화시킨다. 한편으로는 각질층을 침투한 균들은 toll-like receptor (TLR)를 활성화하게 된다. 이후 각질형성세포에서는 여러 케모카인과 TNF-α, IL-1, thymic stromal lymphopoietin (TSLP) 등의 사이토카인들이 분비되어 아토피피부염의 발병을 촉진하고 질환을 악화시킨다[22]. 또한 밀착연접 단백질의 발현이 아토피피부염 환자의 피부에서 정상인에 비하여 현저하게 감소하여 있으며, 최근에는 아토피피부염 환자에서 claudin 1 유전자 (CDN1) 변이가 보고되었고, 헤르페스 바이러스 감염이 쉽게 일어나는 원인이 밀폐연접 단백질 중 claudin-1의 감소 때문이라 설명하고 있다[9,23].
아토피피부염에서 나타나는 비정상적인 면역반응(abnormal immune response) 중 비교적 잘 정리된 것으로는 혈청 IgE 및 알레르겐 감작 증가, 급성 병변에서 Th2 사이토카인 증가, cutaneous lymphocyte-associated antigen (CLA)이 발현된 T세포의 증가, 랑게르한스 세포와 염증성 수지상 표피세포(inflammatory dendritic epidermal cell)에서 FcεRI 발현 증가 등이 있다[24]. 피부장벽이 손상되면 외부 항원 및 균의 침투를 막기 위하여 피부에서는 각질형성세포와 항원전달세포들을 통하여 즉각적인 선천면역반응이 일어나고 후천면역반응도 이어지는데, 아토피피부염 환자에서는 이들 선천면역계와 후천면역계가 비정상적으로 반응하여서 알레르기 염증반응이 발생하는 것으로 설명하고 있다.
피부의 각질형성세포와 항원전달세포는 많은 수의 pattern recognition receptors라고 불리는 선천면역 수용체를 발현하는데 toll like receptors (TLRs)가 가장 널리 알려져 있다[27]. 피부장벽의 손상이나 외부 균에 의하여 TLRs가 자극되면 항균 펩타이드, 사이토카인, 케모카인 등을 분비하고, 더 이상의 외부 균 침투를 막기 위하여 밀폐연접을 강화하게 유도한다[28]. 아토피피부염 환자에서 이런 TLR의 기능이 감소되어 있는 것이 발견되었다. 즉, 아토피피부염 환자에서 나타나는 대표적인 선천면역계 이상은 TLR2 유전자(TLR2)의 과오돌연변이(missense mutation)이다. 이 TLR2 변이를 가진 아토피피부염 환자는 임상적 중증도가 심하고, 혈청 IgE가 높고, 황색포도알균 감염이 증가한다. 한편으로는 Th2 사이토카인에 의해 표피 항균 펩타이드 발현이 감소하는데 이것도 아토피피부염 환자가 세균과 바이러스에 의한 감염에 취약하게 만드는데 관여한다[27].
아토피피부염에서 후천면역반응은 급성기 동안에 일어나는 Th2 사이토카인(IL-4, 13, 31)과 Th22 사이토카인(IL-22)의 증가와 연관된다[28]. 아토피피부염의 병변에는 피부로 돌아오는 수용체(the homing receptor for skin)인 CLA가 발현된 T세포가 증가되어 있다. 손상된 피부장벽을 통하여 외부 항원이 침투하게 되면 수지상 세포가 항원을 인식하여 Th2세포를 활성화시킨다. 활성화된 Th2세포에서 IL-4, 5, 10, 13, 17, 31 등의 사이토카인이 분비되며, IL-4, 13에 의해 B세포가 활성화되어 IgE가 분비된다. IL-5에 의해 호산구가 모여들며, 분비된 IgE가 비만세포와 호염구의 IgE 수용체에 결합한 후 히스타민, 중성 단백질분해효소(neutral protease), 프로스타글란딘 D2, 류코트리엔 C4/D4/E4 등의 다양한 염증매개물질을 분비시켜 아토피피부염의 임상증상을 만든다[29]. Th2 사이토카인인 IL-4, 5, 13 등은 표피분화 단백질인 로리크린(loricrine), 인볼루크린(involucrin), 필라그린 등의 발현을 억제하고 항균 펩타이드 생성도 감소시킨다[30]. 유전학적인 측면에서는 사이토카인 관련 유전자에 변이가 발생하면 여러 Th2 염증반응에 영향을 주어 아토피피부염을 유발할 수 있다. 최근 IL-4, IL-4 수용체, IL-13 등의 유전자 변이와 아토피피부염과의 상관성이 보고되었다[31,32,33].
Th22세포는 IL-22를 생산하며 만성 아토피피부염 병변에서 증가되어 있다. 또한 IL-22 수용체가 각질형성세포에서 높게 발현되며, IL-22의 증가는 표피에서 각질형성세포의 최종분화 결함, 표피의 과증식, 가시세포증식증(acanthosis)을 일으키는 요인이 된다. 따라서 아토피피부염 환자의 피부장벽 손상을 더욱 악화시키는 데에는 Th2세포와 Th22세포가 모두 관여한다. Th17세포는 IL-17과 IL-22를 분비하며 급성기 아토피피부염 환자의 혈액과 피부에서 증가되고, 만성기 아토피피부염에서는 Th2세포에서 분비되는 사이토카인에 의해 IL-17의 분비가 줄어든다[34,35]. Th22세포에서는 IL-22를 분비하여 표피의 증식을 야기하며, 주로 만성기의 아토피피부염에서 높게 나타나고 질병의 활성도와도 관련이 있다[35]. 따라서 급성기 아토피피부염에서는 피부장벽 손상에 의해 외부항원이 쉽게 침투하여 랑게르한스 세포에 의해 감작이 일어나며, Th2와 Th17의 염증반응이 우세하여 관련 사이토카인들에 의해 호산구와 호중구가 모이게 되지만, 만성기 아토피피부염에서는 반복되는 감염, 자극물질, 알레르겐 등의 노출 등에 의해 Th1, Th2, Th22의 염증반응이 우세하게 되어 관련 사이토카인에 의해 표피의 증식 및 각질형성세포 손상 등의 반응이 나타난다. 이 과정에서 IgE의 분비가 증가되며 아토피피부염 병변의 랑게르한스세포와 염증성 수지상 표피세포에서는 FcεRI의 발현이 증가하게 된다.
Th1 및 Th2세포와는 다른 종류의 사이토카인을 생산하여 정상적으로 면역억제 기능을 가지는 조절T (Treg)세포가 아토피피부염 환자의 말초혈액 내에 증가되어 있으나, 초항원(superantigen)인 포도알균 내독소(staphylococcal enterotoxin B)에 의해 이런 면역억제능이 소실된다. 이는 아토피피부염에서 초항원이 T세포를 활성화시켜서 피부의 염증을 더 악화시키는 기전을 제시한다. 그러나 많은 경우에 전신과 피부 면역반응 사이에 차이를 보였듯이 아토피피부염의 피부병변에서는 조절 T세포가 결핍되어 있다[24].
아토피피부염에서 유전 및 여러 환경 요인에 의해 피부장벽이 손상되면 외부로부터 침투한 각종 알레르겐에 쉽게 노출되고, 각질형성세포에서는 IL-7과 유사한 사이토카인인 TSLP를 분비한다. TSLP는 알레르기 염증반응의 'master switch'라고도 불리는데 비만세포, 호염구, 호산구 등 피부염증을 일으키는 중요한 세포에 영향을 주고, 선천면역 및 후천면역 반응 모두에 영향을 주며, Th2 면역반응을 유도하여 아토피피부염의 병인에 매우 중요한 요소로 인식되고 있다[36]. 아토피피부염 환자의 피부와 혈액에서는 TSLP가 증가되는데, 알레르겐에 의해 각질형성세포의 PAR-2 발현이 증가되면 표피에서의 TSLP 분비가 증가되고 이로 인하여 Th2 염증반응이 우세하게 일어나서 아토피피부염의 임상양상으로 나타난다[37] (Figure 3). 한편으로는 TSLP 유전자에 돌연변이가 있을 경우에 TSLP 분비가 더 증가되고 Th2 면역반응이 더 심하게 나타난다[38,39]. 또한 알레르기 행진에서 아토피피부염 환자가 천식이나 알레르기 비염으로 진행하는데 중요한 매개자로 TSLP가 거론되고 있어서 TSLP는 아토피피부염의 발생을 예방하는 타깃인 동시에 알레르기 행진으로의 진행을 막는 치료적 타깃으로 생각되고 있다[40].
아토피피부염은 선천적으로 피부장벽의 결함뿐만이 아니라 면역체계의 이상반응이나 결함도 가지고 있는 사람들에게 더욱 흔하게 나타난다. 즉, 피부장벽 기능 이상과 선천면역 및 후천면역 체계의 비정상적인 반응이 상호 복합적으로 아토피피부염의 병인으로 작용 하므로(Figure 4), 전체적인 병인을 아우르는 관리 및 치료법의 선택과 새로운 치료제의 개발이 필요할 것이다.
Acknowledgement
This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korean government (NRF-2012R1A1A4A01013614).
References
1. Yu JS, Lee CJ, Lee HS, Kim J, Han Y, Ahn K, Lee SI. Prevalence of atopic dermatitis in Korea: analysis by using national statistics. J Korean Med Sci 2012;27:681-685.




2. Palmer CN, Irvine AD, Terron-Kwiatkowski A, Zhao Y, Liao H, Lee SP, Goudie DR, Sandilands A, Campbell LE, Smith FJ, O'Regan GM, Watson RM, Cecil JE, Bale SJ, Compton JG, DiGiovanna JJ, Fleckman P, Lewis-Jones S, Arseculeratne G, Ser-geant A, Munro CS, El Houate B, McElreavey K, Halkjaer LB, Bisgaard H, Mukhopadhyay S, McLean WH. Common loss-of-function variants of the epidermal barrier protein filaggrin are a major predisposing factor for atopic dermatitis. Nat Genet 2006;38:441-446.



3. Elias PM, Steinhoff M. "Outside-to-inside" (and now back to "outside") pathogenic mechanisms in atopic dermatitis. J Invest Dermatol 2008;128:1067-1070.



4. Choi EH, Yoon NY, Kim DH, Jung M. Barrier related biomarkers of atopic dermatitis. J Skin Barrier Res 2013;15:13-23.
5. Irvine AD, McLean WH. Breaking the (un)sound barrier: filaggrin is a major gene for atopic dermatitis. J Invest Dermatol 2006;126:1200-1202.


6. Melnik B, Hollmann J, Plewig G. Decreased stratum corneum ceramides in atopic individuals: a pathobiochemical factor in xerosis? Br J Dermatol 1988;119:547-549.


7. Ong PY, Ohtake T, Brandt C, Strickland I, Boguniewicz M, Ganz T, Gallo RL, Leung DY. Endogenous antimicrobial peptides and skin infections in atopic dermatitis. N Engl J Med 2002;347:1151-1160.


8. Kato A, Fukai K, Oiso N, Hosomi N, Murakami T, Ishii M. Association of SPINK5 gene polymorphisms with atopic dermatitis in the Japanese population. Br J Dermatol 2003;148:665-669.


9. De Benedetto A, Rafaels NM, McGirt LY, Ivanov AI, Georas SN, Cheadle C, Berger AE, Zhang K, Vidyasagar S, Yoshida T, Boguniewicz M, Hata T, Schneider LC, Hanifin JM, Gallo RL, Novak N, Weidinger S, Beaty TH, Leung DY, Barnes KC, Beck LA. Tight junction defects in patients with atopic dermatitis. J Allergy Clin Immunol 2011;127:773-786e7.


10. Seguchi T, Cui CY, Kusuda S, Takahashi M, Aisu K, Tezuka T. Decreased expression of filaggrin in atopic skin. Arch Dermatol Res 1996;288:442-446.



11. Choi EH, Yoon NY. Skin barrier related pathogenesis of atopic dermatitis. J Skin Barrier Res 2012;14:70-76.

12. Morar N, Cookson WO, Harper JI, Moffatt MF. Filaggrin mutations in children with severe atopic dermatitis. J Invest Dermatol 2007;127:1667-1672.


13. Nomura T, Akiyama M, Sandilands A, Nemoto-Hasebe I, Sakai K, Nagasaki A, Ota M, Hata H, Evans AT, Palmer CN, Shimizu H, McLean WH. Specific filaggrin mutations cause ichthyosis vulgaris and are significantly associated with atopic dermatitis in Japan. J Invest Dermatol 2008;128:1436-1441.


14. Li M, Liu Q, Liu J, Cheng R, Zhang H, Xue H, Bao Y, Yao Z. Mutations analysis in filaggrin gene in northern China patients with atopic dermatitis. J Eur Acad Dermatol Venereol 2013;27:169-174.


15. Greisenegger E, Novak N, Maintz L, Bieber T, Zimprich F, Haubenberger D, Gleiss A, Stingl G, Kopp T, Zimprich A. Analysis of four prevalent filaggrin mutations (R501X, 2282del4, R2447X and S3247X) in Austrian and German patients with atopic dermatitis. J Eur Acad Dermatol Venereol 2010;24:607-610.


16. Barker JN, Palmer CN, Zhao Y, Liao H, Hull PR, Lee SP, Allen MH, Meggitt SJ, Reynolds NJ, Trembath RC, McLean WH. Null mutations in the filaggrin gene (FLG) determine major susceptibility to early-onset atopic dermatitis that persists into adulthood. J Invest Dermatol 2007;127:564-567.


17. Kezic S, O'Regan GM, Yau N, Sandilands A, Chen H, Campbell LE, Kroboth K, Watson R, Rowland M, McLean WH, Irvine AD. Levels of filaggrin degradation products are influenced by both filaggrin genotype and atopic dermatitis severity. Allergy 2011;66:934-940.



18. Imokawa G, Abe A, Jin K, Higaki Y, Kawashima M, Hidano A. Decreased level of ceramides in stratum corneum of atopic dermatitis: an etiologic factor in atopic dry skin? J Invest Dermatol 1991;96:523-526.


19. Jeong SK, Kim HJ, Youm JK, Ahn SK, Choi EH, Sohn MH, Kim KE, Hong JH, Shin DM, Lee SH. Mite and cockroach allergens activate protease-activated receptor 2 and delay epidermal permeability barrier recovery. J Invest Dermatol 2008;128:1930-1939.


20. Zhao LP, Di Z, Zhang L, Wang L, Ma L, Lv Y, Hong Y, Wei H, Chen HD, Gao XH. Association of SPINK5 gene polymorphisms with atopic dermatitis in Northeast China. J Eur Acad Dermatol Venereol 2012;26:572-577.


21. Baek JH, Lee SE, Choi KJ, Choi EH, Lee SH. Acute modulations in stratum corneum permeability barrier function affect claudin expression and epidermal tight junction function via changes of epidermal calcium gradient. Yonsei Med J 2013;54:523-528.



22. Kubo A, Nagao K, Amagai M. Epidermal barrier dysfunction and cutaneous sensitization in atopic diseases. J Clin Invest 2012;122:440-447.



23. De Benedetto A, Slifka MK, Rafaels NM, Kuo IH, Georas SN, Boguniewicz M, Hata T, Schneider LC, Hanifin JM, Gallo RL, Johnson DC, Barnes KC, Leung DY, Beck LA. Reductions in claudin-1 may enhance susceptibility to herpes simplex virus 1 infections in atopic dermatitis. J Allergy Clin Immunol 2011;128:242-246e5.



24. Boguniewicz M, Leung DY. Recent insights into atopic dermatitis and implications for management of infectious complications. J Allergy Clin Immunol 2010;125:4-13.



25. Valenta R, Natter S, Seiberler S, Roschanak M, Mothes N, Mahler V, Eibensteiner P. Autoallergy: a pathogenetic factor in atopic dermatitis? Curr Probl Dermatol 1999;28:45-50.


26. Liu FT, Goodarzi H, Chen HY. IgE, mast cells, and eosinophils in atopic dermatitis. Clin Rev Allergy Immunol 2011;41:298-310.



27. Kuo IH, Yoshida T, De Benedetto A, Beck LA. The cutaneous innate immune response in patients with atopic dermatitis. J Allergy Clin Immunol 2013;131:266-278.


28. Leung DY. New insights into atopic dermatitis: role of skin barrier and immune dysregulation. Allergol Int 2013;62:151-161.



29. Mamessier E, Magnan A. Cytokines in atopic diseases: revisiting the Th2 dogma. Eur J Dermatol 2006;16:103-113.

30. Howell MD, Boguniewicz M, Pastore S, Novak N, Bieber T, Girolomoni G, Leung DY. Mechanism of HBD-3 deficiency in atopic dermatitis. Clin Immunol 2006;121:332-338.


31. He JQ, Chan-Yeung M, Becker AB, Dimich-Ward H, Ferguson AC, Manfreda J, Watson WT, Sandford AJ. Genetic variants of the IL13 and IL4 genes and atopic diseases in at-risk children. Genes Immun 2003;4:385-389.



32. Beghe B, Barton S, Rorke S, Peng Q, Sayers I, Gaunt T, Keith TP, Clough JB, Holgate ST, Holloway JW. Polymorphisms in the interleukin-4 and interleukin-4 receptor alpha chain genes confer susceptibility to asthma and atopy in a Caucasian population. Clin Exp Allergy 2003;33:1111-1117.

33. Hummelshoj T, Bodtger U, Datta P, Malling HJ, Oturai A, Poulsen LK, Ryder LP, Sorensen PS, Svejgaard E, Svejgaard A. Association between an interleukin-13 promoter polymorphism and atopy. Eur J Immunogenet 2003;30:355-359.



34. Koga C, Kabashima K, Shiraishi N, Kobayashi M, Tokura Y. Possible pathogenic role of Th17 cells for atopic dermatitis. J Invest Dermatol 2008;128:2625-2630.


35. Nograles KE, Zaba LC, Shemer A, Fuentes-Duculan J, Cardinale I, Kikuchi T, Ramon M, Bergman R, Krueger JG, Guttman-Yassky E. IL-22-producing "T22" T cells account for upregulated IL-22 in atopic dermatitis despite reduced IL-17-producing TH17 T cells. J Allergy Clin Immunol 2009;123:1244-1252e2.



36. Ziegler SF, Artis D. Sensing the outside world: TSLP regulates barrier immunity. Nat Immunol 2010;11:289-293.




37. Lee EB, Kim KW, Hong JY, Jee HM, Sohn MH, Kim KE. Increased serum thymic stromal lymphopoietin in children with atopic dermatitis. Pediatr Allergy Immunol 2010;21:e457-e460.


38. Wu WH, Park CO, Oh SH, Kim HJ, Kwon YS, Bae BG, Noh JY, Lee KH. Thymic stromal lymphopoietin-activated invariant natural killer T cells trigger an innate allergic immune response in atopic dermatitis. J Allergy Clin Immunol 2010;126:290-299e4.


39. Harada M, Hirota T, Jodo AI, Hitomi Y, Sakashita M, Tsunoda T, Miyagawa T, Doi S, Kameda M, Fujita K, Miyatake A, Enomoto T, Noguchi E, Masuko H, Sakamoto T, Hizawa N, Suzuki Y, Yoshihara S, Adachi M, Ebisawa M, Saito H, Matsumoto K, Nakajima T, Mathias RA, Rafaels N, Barnes KC, Himes BE, Duan QL, Tantisira KG, Weiss ST, Nakamura Y, Ziegler SF, Tamari M. Thymic stromal lymphopoietin gene promoter polymorphisms are associated with susceptibility to bronchial asthma. Am J Respir Cell Mol Biol 2011;44:787-793.


Figure 1
Life cycle of filaggrin. Filaggrin exists as profilaggrin within keratohyaline granules in the granular layer of epidermis. It is degraded to filaggrin during the terminal differentiation process. Then, going up to the upper part of stratum corneum (SC), filaggrin degrades into amino acids and plays a crucial role in maintaining SC hydration and pH by forming natural moisturizing factor including pyrrolidone carboxylic acid or urocanic acid (From Choi EH, et al. J Skin Barrier Res 2013;15:13-23, with permission from Korean Society for Skin Barrier Research) [4].
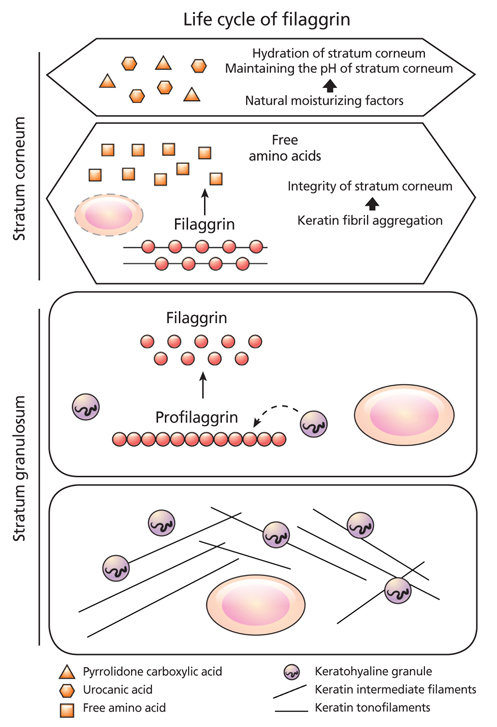
Figure 2
Two main pathogenesis of dry skin in atopic dermatitis. First pathway is a reduced natural moisturizing factor (NMF) in the stratum corneum (SC) due to filaggrin deficiency which caused by inherited FLG mutation or lesional Th2 inflammation. Second is a decreased SC hydration as well as increased transepidermal water loss (TEWL) and epidermal proliferation due to disrupted SC intercellular lamellae which caused by deficiency of ceramide (esp, ceramide 1 having long chain) in SC. KC, keratinocytes.
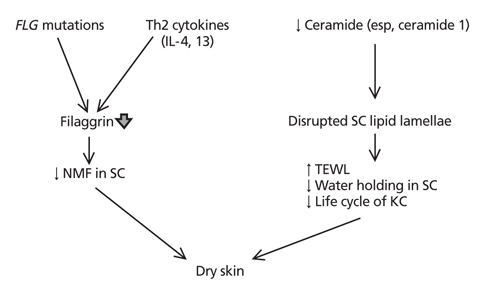
Figure 3
Complex interaction among skin pH, filaggrin, serine protease and serine protease inhibitor in skin inflammation of atopic dermatitis (AD) patients. Acute barrier disruption in non-atopy healthy people induces serine protease (SP) activation transiently, but inhibited immediately with serine protease inhibitor (SPI), therefore reaches to barrier recovery (top blue dotted box). On the other hand, in the case of AD patients with SPINK5 or FLG gene mutation, uncontrolled SP activity could induce the activation of protease-activated receptor-2 (PAR-2), in turn thymic stromal lymphopoietin (TSLP) followed by aggravation cycle to skin allergic inflammation (bottom red dotted box). LB, lamellar body, LC, Langerhans cell.
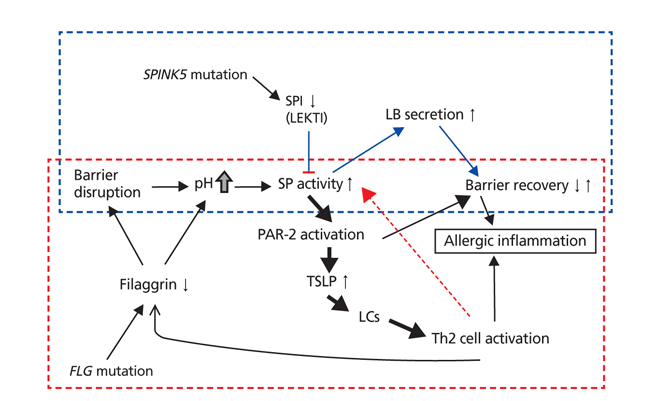
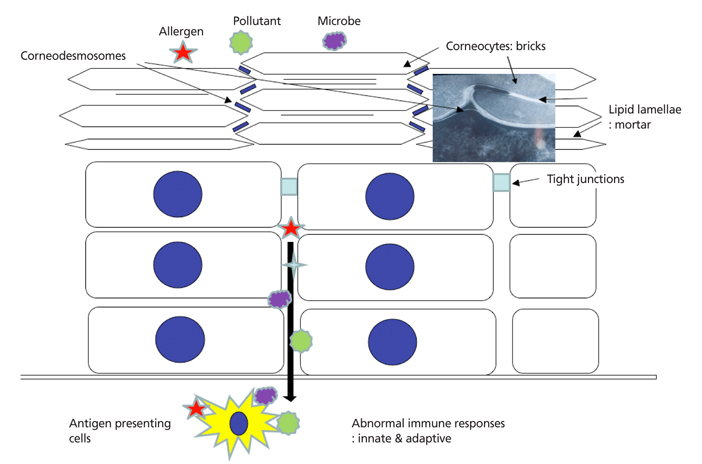
Figure 4
Combined pathogenesis of atopic dermatitis including dysfunctions of skin barrier and immune response. Skin barrier is composed of corneocytes (brick), intercellular lipid lamellae (mortar) and corneodesmosomes (rivet) in stratum corneum and tight junctions in upper stratum granulosum. If these structures have inherited or acquired defect, exogenous allergens, microbes, or pollutants can easily penetrate skin barrier. And then abnormal responses of innate immune and adaptive system induce allergic inflammation in the patients with atopic dermatitis.
- TOOLS
-
METRICS

-
Related articles in
J Korean Med Assoc -
Epidemiology and pathogenesis of scabies2023 December;66(12)
Therapeutic approach to atopic dermatitis in children2022 October;65(10)
Diagnosis of varicose veins2022 April;65(4)
Pathophysiology of neuropathic pain2021 July;64(7)
Pathogenesis and prevention of pressure ulcer2021 January;64(1)






